Exercises
Targeted exercises
Implementing linear interpolation using numpy.interp
Exercise 1
Consider a Gaussian distribution with $x_0 = 0$, $\sigma = 2$, and amplitude of 1.
- Create 1000 $x$-values between -10 and 10, and use this to create y-values with the above Gaussian parameters. This will represent the ’true’ Guassian shape.
- Repeat the above, but with 5, 6, 7, 8, 9, 10, 11, 12 points, each from -10 and 10. These will represent sparsely sampled points.
- For each set of points generated in the second step, perform a linear interpolation using
numpy.interp(). This accepts three positional arguments: an array of $x$-values you wisht to generate interpolated points for, the original $x$-values, and the original $y$ values used for interpolation. - Make a Plotly plot holding a separate subplot for each of the eight sparse arrays. On this plot, have the ’true’ Gaussian shape, the sparsely sampled points, and the interpolated values.
- Comment on how the interpolation improves with sample points.
- Comment on what regions of the Gaussian seem to be reproduced well, and which are not.
import numpy as np
from plotly.subplots import make_subplots
def gaussian(x, x0, sigma, amplitude):
return amplitude * np.exp(-((x - x0) ** 2) / (2 * sigma**2))
fig = make_subplots(rows = 8, cols = 1)
ultra_fine_x = np.linspace(-10,10,1000)
ultra_fine_y = gaussian(ultra_fine_x, x0=0, sigma=2, amplitude=1)
for i, n in enumerate(range(5, 13)):
fig.add_scatter(x = ultra_fine_x, y = ultra_fine_y, line = dict(color = "grey", width = 10),
showlegend = False,
row = i+1, col = 1)
x_coarse = np.linspace(-10, 10, n)
y_coarse = gaussian(x_coarse, x0=0, sigma=2, amplitude=1)
fig.add_scatter(x = x_coarse, y = y_coarse,
mode = "markers",
showlegend = False,
marker = dict(color = "red", size = 15),
row = i+1, col = 1)
lin_y = np.interp(ultra_fine_x, x_coarse, y_coarse)
fig.add_scatter(x = ultra_fine_x, y = lin_y, line = dict(color = "orange"),
showlegend = False,
row = i+1, col = 1)
fig.add_annotation(text = f"n={n}",
showarrow = False,
x = -9, y = 0.2,
row = i+1, col = 1
)
fig.update_layout(template = "simple_white", height = 1200, width = 600)
fig.show("png")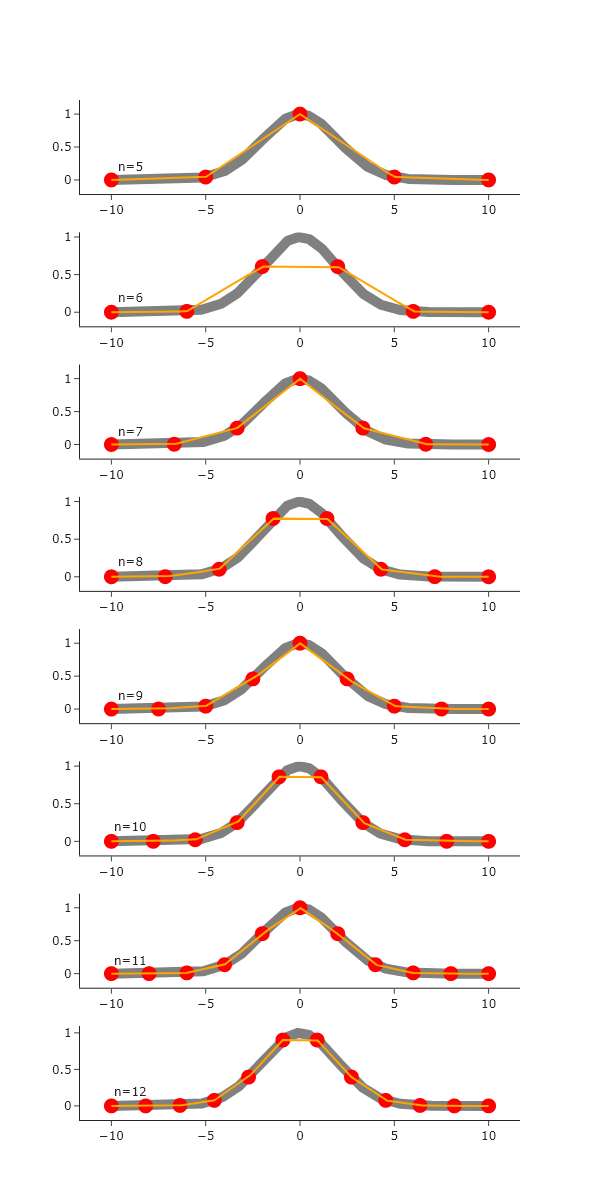
 As the number of sparse points increases, the interpolation better agrees with the ’true’ Gaussian, which is expected. The linear interpolation seems to handle regions with out curvature well, but struggles where there is significant curvature.
As the number of sparse points increases, the interpolation better agrees with the ’true’ Gaussian, which is expected. The linear interpolation seems to handle regions with out curvature well, but struggles where there is significant curvature.
Implementing cubic spline interpolation using scipy.interpolate.CubicSpline
Exercise 1
Repeat Exercise 0, but use a cubic spline for interpolation.
In addition to answering all parts from Exercise 0, comment on the differences between the results from the linear interpolation, versus the cubic interpolation.
import numpy as np
from plotly.subplots import make_subplots
import scipy.interpolate as spi
def gaussian(x, x0, sigma, amplitude):
return amplitude * np.exp(-((x - x0) ** 2) / (2 * sigma**2))
fig = make_subplots(rows = 8, cols = 1)
ultra_fine_x = np.linspace(-10,10,1000)
ultra_fine_y = gaussian(ultra_fine_x, x0=0, sigma=2, amplitude=1)
for i, n in enumerate(range(5, 13)):
fig.add_scatter(x = ultra_fine_x, y = ultra_fine_y, line = dict(color = "grey", width = 10),
showlegend = False,
row = i+1, col = 1)
x_coarse = np.linspace(-10, 10, n)
y_coarse = gaussian(x_coarse, x0=0, sigma=2, amplitude=1)
fig.add_scatter(x = x_coarse, y = y_coarse,
mode = "markers",
showlegend = False,
marker = dict(color = "red", size = 15),
row = i+1, col = 1)
y_cs = spi.CubicSpline(x_coarse, y_coarse)
y_cubic = y_cs(ultra_fine_x)
fig.add_scatter(x = ultra_fine_x, y = y_cubic, line = dict(color = "orange"),
showlegend = False,
row = i+1, col = 1)
fig.add_annotation(text = f"n={n}",
showarrow = False,
x = -9, y = 0.2,
row = i+1, col = 1
)
fig.update_layout(template = "simple_white", height = 1200, width = 600)
fig.show("png")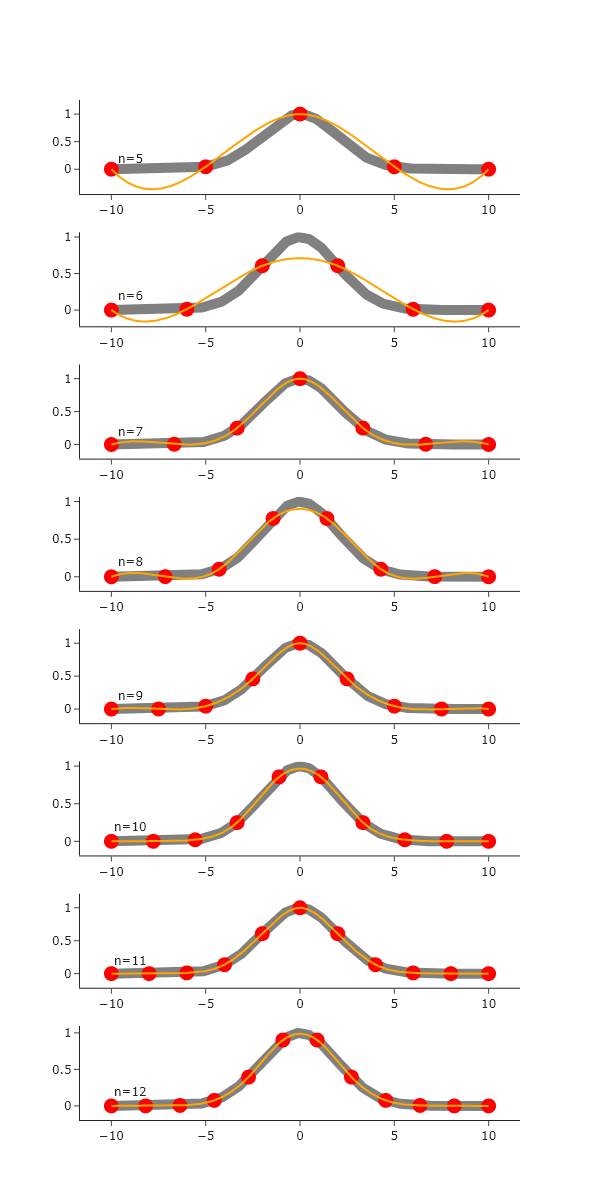
 Similar to the linear interpolation, the agreement of the interpolation with the ’true’ Gaussian value increases as the number of sparse points increases. However, the cubic interpolation handles regions of curvature well, and struggles with regions that do not have curvature. The overall agreement also seems to be better for cubic, rather than linear, once the number of points reaches ~9. Thus, as the sampling increases, the cubic interpolation is the clear winner.
Similar to the linear interpolation, the agreement of the interpolation with the ’true’ Gaussian value increases as the number of sparse points increases. However, the cubic interpolation handles regions of curvature well, and struggles with regions that do not have curvature. The overall agreement also seems to be better for cubic, rather than linear, once the number of points reaches ~9. Thus, as the sampling increases, the cubic interpolation is the clear winner.
Comprehensive
Exercise 2
Modify the final code from Chapter 8 to interpolate the kinetic traces so that they have evenly spaced points. Use both linear interpolation and cubic splines. Do you notice any obvious inaccuracies for either method?
'''
A code to numerically solve a reaction mechanism that is irreversible but
has two intermediates using solve_ivp from scipy.integrate
Written by: Ben Lear and Chris Johnson (authors@codechembook.com)
v1.0.0 - 250217 - initial version
v1.0.1 - 250406 - changed so that the spacing along the final t-values is even
'''
import scipy.integrate as spi
from codechembook.quickPlots import quickScatter
import scipy.interpolate as spint
import numpy as np
# First we need to define a function containing our differential rate laws.
def TwoIntermediatesDE(t, y, k):
'''
Differential rate laws for a catalytic reaction with two intermediates:
A + cat ->(k1) X
X + B ->(k2) Y
Y ->(k3) C + cat
REQUIRED PARAMETERS:
t (float): the cutrent time in the simulation.
Not explicitly used but needed by solve_ivp
y (list of float): the current concentration
k (list of float): the rate
RETURNS:
(list of float): rate of change of concentration in the order of y
'''
A, B, cat, X, Y, C = y # unpack concentrations to convenient variables
k1, k2, k3 = k # unpack rates
dAdt = -k1 * A * cat
dBdt = -k2 * X * B
dcatdt = -k1 * A * cat + k3 * Y
dXdt = k1 * A * cat - k2 * X * B
dYdt = k2 * X * B - k3 * Y
dCdt = k3 * Y
return [dAdt, dBdt, dcatdt, dXdt, dYdt, dCdt]
# Set up initial conditions and simulation parameters
y0 = [1.0, 1.0, 0.2, 0.0, 0.0, 0.0] # concetrations (mM) [A, B, cat, X, Y, C]
k = [5e1, 1e1, 5e0] # rate (1/s)
time = [0, 10] # simulation start and end times (s)
# Invoke solve_ivp and store the result object
solution = spi.solve_ivp(TwoIntermediatesDE, time, y0, args = [k])
t_int = np.linspace(0,10,1000) # set up the even spacing along x
y_ints = []
for y in solution.y:
y_cs = spint.CubicSpline(solution.t, y)
y_ints.append(y_cs(t_int))
# Plot the results
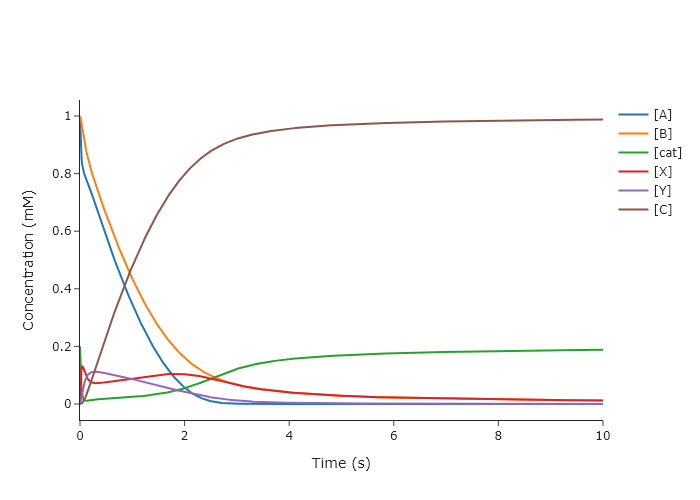
quickScatter(x = t_int, # need a list of six identical time axes
y = y_ints,
name = ['[A]', '[B]', '[cat]', '[X]', '[Y]', '[C]'],
xlabel = 'Time (s)',
ylabel = 'Concentration (mM)',
output = 'png')